| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-09-30 23:26:02 UTC |
|---|
| Update Date | 2020-05-21 16:28:41 UTC |
|---|
| BMDB ID | BMDB0006792 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
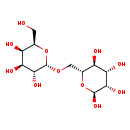
| Common Name | Epimelibiose |
|---|
| Description | Melibiose, also known as isomaltose, belongs to the class of organic compounds known as o-glycosyl compounds. These are glycoside in which a sugar group is bonded through one carbon to another group via a O-glycosidic bond. Melibiose is possibly soluble (in water) and an extremely weak basic (essentially neutral) compound (based on its pKa). Melibiose exists in all living species, ranging from bacteria to humans. Melibiose participates in a number of enzymatic reactions, within cattle. In particular, Melibiose and D-galactose can be biosynthesized from Alpha-lactose; which is mediated by the enzyme Alpha-galactosidase a. In addition, Melibiose can be converted into D-galactose through its interaction with the enzyme Alpha-galactosidase a. In cattle, melibiose is involved in the metabolic pathway called the galactose metabolism pathway. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 6-O-alpha-D-Galactopyranosyl-D-mannose | HMDB | | 6-O-alpha-D-Galactopyranosyl-alpha-D-mannopyranose | HMDB | | 6-O-Α-D-galactopyranosyl-D-mannose | HMDB | | 6-O-Α-D-galactopyranosyl-α-D-mannopyranose | HMDB | | D-Epimelibiose | HMDB | | alpha-Epimelibiose | HMDB | | Α-epimelibiose | HMDB | | Epimelibiose | HMDB |
|
|---|
| Chemical Formula | C12H22O11 |
|---|
| Average Molecular Weight | 342.297 |
|---|
| Monoisotopic Molecular Weight | 342.116211528 |
|---|
| IUPAC Name | (2S,3S,4S,5S,6R)-6-({[(2S,3R,4S,5R,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}methyl)oxane-2,3,4,5-tetrol |
|---|
| Traditional Name | (2S,3S,4S,5S,6R)-6-({[(2S,3R,4S,5R,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}methyl)oxane-2,3,4,5-tetrol |
|---|
| CAS Registry Number | 585-99-9 |
|---|
| SMILES | OC[C@H]1O[C@H](OC[C@H]2O[C@H](O)[C@@H](O)[C@@H](O)[C@@H]2O)[C@H](O)[C@@H](O)[C@H]1O |
|---|
| InChI Identifier | InChI=1S/C12H22O11/c13-1-3-5(14)8(17)10(19)12(23-3)21-2-4-6(15)7(16)9(18)11(20)22-4/h3-20H,1-2H2/t3-,4-,5+,6-,7+,8+,9+,10-,11+,12+/m1/s1 |
|---|
| InChI Key | DLRVVLDZNNYCBX-LGHYCPABSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as o-glycosyl compounds. These are glycoside in which a sugar group is bonded through one carbon to another group via a O-glycosidic bond. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | O-glycosyl compounds |
|---|
| Alternative Parents | |
|---|
| Substituents | - O-glycosyl compound
- Disaccharide
- Oxane
- Secondary alcohol
- Hemiacetal
- Oxacycle
- Organoheterocyclic compound
- Polyol
- Acetal
- Hydrocarbon derivative
- Primary alcohol
- Alcohol
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Ontology |
|---|
| Status | Expected but not Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-002f-0209000000-7959f78227a7a5ad2e03 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03fv-4954000000-b770df15d17722e22d68 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0kbv-9340000000-ecfae98627db56099495 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0029000000-ed9511fe35d9af9ada39 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0pi3-6294000000-cffe78babaec65350402 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9100000000-341f45dca0cdd3356a31 | View in MoNA |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
|
|---|